| Weight | 1 lbs |
|---|---|
| Dimensions | 9 × 5 × 2 in |
| host | mouse |
| isotype | IgG2b |
| clonality | monoclonal |
| concentration | concentrate, predilute |
| applications | IHC |
| reactivity | human |
| available size | 0.1 mL, 0.5 mL, 1 mL concentrated, 7 mL prediluted |
mouse anti-Beta-Catenin monoclonal antibody (ZM13) 6033
Price range: $160.00 through $528.00
Antibody summary
- Mouse monoclonal to Beta-Catenin
- Suitable for: Immunohistochemistry (formalin-fixed, paraffin-embedded tissues)
- Reacts with: Human
- Isotype:IgG2b
- Control: Fibromatosis or transitional cell carcinoma
- Visualization: Membrane, cytoplasmic or nuclear
- 0.1, 0.5, 1.0 mL concentrated, 7 mL prediluted
mouse anti-Beta-Catenin monoclonal antibody ZM13 6033
| target relevance |
|---|
| Protein names Catenin beta-1 (Beta-catenin) |
| Gene names CTNNB1,CTNNB1 CTNNB OK/SW-cl.35 PRO2286 |
| Protein family Beta-catenin family |
| Mass 85497Da |
| Function FUNCTION: Key downstream component of the canonical Wnt signaling pathway (PubMed:17524503, PubMed:18077326, PubMed:18086858, PubMed:18957423, PubMed:21262353, PubMed:22155184, PubMed:22647378, PubMed:22699938). In the absence of Wnt, forms a complex with AXIN1, AXIN2, APC, CSNK1A1 and GSK3B that promotes phosphorylation on N-terminal Ser and Thr residues and ubiquitination of CTNNB1 via BTRC and its subsequent degradation by the proteasome (PubMed:17524503, PubMed:18077326, PubMed:18086858, PubMed:18957423, PubMed:21262353, PubMed:22155184, PubMed:22647378, PubMed:22699938). In the presence of Wnt ligand, CTNNB1 is not ubiquitinated and accumulates in the nucleus, where it acts as a coactivator for transcription factors of the TCF/LEF family, leading to activate Wnt responsive genes (PubMed:17524503, PubMed:18077326, PubMed:18086858, PubMed:18957423, PubMed:21262353, PubMed:22155184, PubMed:22647378, PubMed:22699938). Also acts as a coactivator for other transcription factors, such as NR5A2 (PubMed:22187462). Promotes epithelial to mesenchymal transition/mesenchymal to epithelial transition (EMT/MET) via driving transcription of CTNNB1/TCF-target genes (PubMed:29910125). Involved in the regulation of cell adhesion, as component of an E-cadherin:catenin adhesion complex (By similarity). Acts as a negative regulator of centrosome cohesion (PubMed:18086858). Involved in the CDK2/PTPN6/CTNNB1/CEACAM1 pathway of insulin internalization (PubMed:21262353). Blocks anoikis of malignant kidney and intestinal epithelial cells and promotes their anchorage-independent growth by down-regulating DAPK2 (PubMed:18957423). Disrupts PML function and PML-NB formation by inhibiting RANBP2-mediated sumoylation of PML (PubMed:22155184). Promotes neurogenesis by maintaining sympathetic neuroblasts within the cell cycle (By similarity). Involved in chondrocyte differentiation via interaction with SOX9: SOX9-binding competes with the binding sites of TCF/LEF within CTNNB1, thereby inhibiting the Wnt signaling (By similarity). Acts as a positive regulator of odontoblast differentiation during mesenchymal tooth germ formation, via promoting the transcription of differentiation factors such as LEF1, BMP2 and BMP4 (By similarity). Activity is repressed in a MSX1-mediated manner at the bell stage of mesenchymal tooth germ formation which prevents premature differentiation of odontoblasts (By similarity). {ECO:0000250|UniProtKB:Q02248, ECO:0000269|PubMed:17524503, ECO:0000269|PubMed:18077326, ECO:0000269|PubMed:18086858, ECO:0000269|PubMed:18957423, ECO:0000269|PubMed:21262353, ECO:0000269|PubMed:22155184, ECO:0000269|PubMed:22187462, ECO:0000269|PubMed:22647378, ECO:0000269|PubMed:22699938, ECO:0000269|PubMed:29910125}. |
| Subellular location SUBCELLULAR LOCATION: Cytoplasm {ECO:0000269|PubMed:22294297, ECO:0000269|PubMed:25183871, ECO:0000269|PubMed:29739711, ECO:0000269|PubMed:29910125, ECO:0000269|PubMed:31801859}. Nucleus {ECO:0000269|PubMed:21118991, ECO:0000269|PubMed:24342833, ECO:0000269|PubMed:25183871, ECO:0000269|PubMed:28829046, ECO:0000269|PubMed:29367600, ECO:0000269|PubMed:29739711, ECO:0000269|PubMed:29910125, ECO:0000269|PubMed:31801859}. Cytoplasm, cytoskeleton {ECO:0000250|UniProtKB:B6V8E6}. Cell junction, adherens junction {ECO:0000269|PubMed:10725230, ECO:0000269|PubMed:22294297}. Cell junction {ECO:0000250|UniProtKB:B6V8E6}. Cell membrane {ECO:0000269|PubMed:11790773, ECO:0000269|PubMed:24342833}. Cytoplasm, cytoskeleton, microtubule organizing center, centrosome. Cytoplasm, cytoskeleton, spindle pole. Synapse {ECO:0000250|UniProtKB:Q02248}. Cytoplasm, cytoskeleton, cilium basal body {ECO:0000250|UniProtKB:Q02248}. Note=Colocalized with RAPGEF2 and TJP1 at cell-cell contacts (By similarity). Cytoplasmic when it is un-stable (highly phosphorylated) or bound to CDH1. Translocates to the nucleus when it is stabilized (low level of phosphorylation). Interaction with GLIS2 and MUC1 promotes nuclear translocation. Interaction with EMD inhibits nuclear localization. The majority of CTNNB1 is localized to the cell membrane. In interphase, colocalizes with CROCC between CEP250 puncta at the proximal end of centrioles, and this localization is dependent on CROCC and CEP250. In mitosis, when NEK2 activity increases, it localizes to centrosomes at spindle poles independent of CROCC. Colocalizes with CDK5 in the cell-cell contacts and plasma membrane of undifferentiated and differentiated neuroblastoma cells. Interaction with FAM53B promotes translocation to the nucleus (PubMed:25183871). Translocates to the nucleus in the presence of SNAIL1 (By similarity). Ca(2+)-mediated localization to the cell membrane in dental epithelial cells is inhibited via WNT3A (By similarity). Localizes to cell-cell contacts as keratinocyte differentiation progresses (By similarity). {ECO:0000250|UniProtKB:B6V8E6, ECO:0000250|UniProtKB:Q02248, ECO:0000269|PubMed:25183871}. |
| Tissues TISSUE SPECIFICITY: Expressed in several hair follicle cell types: basal and peripheral matrix cells, and cells of the outer and inner root sheaths. Expressed in colon. Present in cortical neurons (at protein level). Expressed in breast cancer tissues (at protein level) (PubMed:29367600). {ECO:0000269|PubMed:11703283, ECO:0000269|PubMed:17009320, ECO:0000269|PubMed:17289029, ECO:0000269|PubMed:29367600}. |
| Structure SUBUNIT: Two separate complex-associated pools are found in the cytoplasm. The majority is present as component of an E-cadherin:catenin adhesion complex composed of at least E-cadherin/CDH1 and beta-catenin/CTNNB1, and possibly alpha-catenin/CTNNA1; the complex is located to adherens junctions. The stable association of CTNNA1 is controversial as CTNNA1 was shown not to bind to F-actin when assembled in the complex. Alternatively, the CTNNA1-containing complex may be linked to F-actin by other proteins such as LIMA1. Another cytoplasmic pool is part of a large complex containing AXIN1, AXIN2, APC, CSNK1A1 and GSK3B that promotes phosphorylation on N-terminal Ser and Thr residues and ubiquitination of CTNNB1 via BTRC and its subsequent degradation by the proteasome. Wnt-dependent activation of DVL antagonizes the action of GSK3B. When GSK3B activity is inhibited the complex dissociates, CTNNB1 is dephosphorylated and is no longer targeted for destruction. The stabilized protein translocates to the nucleus, where it binds TCF/LEF-1 family members, BCL9, BCL9L and possibly also RUVBL1 and CHD8. Binds CTNNBIP and EP300. CTNNB1 forms a ternary complex with LEF1 and EP300 that is disrupted by CTNNBIP1 binding. Interacts with TAX1BP3 (via the PDZ domain); this interaction inhibits the transcriptional activity of CTNNB1. Interacts with AJAP1, BAIAP1, CARM1, CTNNA3, CXADR and PCDH11Y. Binds NHERF1. Interacts with GLIS2 and MUC1. Interacts with SLC30A9. Interacts with XIRP1. Interacts directly with AXIN1; the interaction is regulated by CDK2 phosphorylation of AXIN1. Interacts with SCRIB. Interacts with RAPGEF2. Interacts with PTPRU (via the cytoplasmic juxtamembrane domain). Interacts with EMD. Interacts with TNIK and TCF7L2. Interacts with SESTD1 and TRPC4. Interacts with CAV1. Interacts with TRPV4. The TRPV4 and CTNNB1 complex can interact with CDH1. Interacts with VCL. Interacts with PTPRJ. Interacts with PKT7 and CDK2. Interacts with FAT1 (via the cytoplasmic domain). Interacts with NANOS1 and NDRG2. Interacts with isoform 1 of NEK2. Interacts with both isoform 1 and isoform 2 of CDK5. Interacts with PTK6. Interacts with SOX7; this interaction may lead to proteasomal degradation of active CTNNB1 and thus inhibition of Wnt/beta-catenin-stimulated transcription. Identified in a complex with HINT1 and MITF. Interacts with FHIT. The CTNNB1 and TCF7L2/TCF4 complex interacts with PML (isoform PML-4). Interacts with FERMT2. Identified in a complex with TCF7L2/TCF4 and FERMT2 (PubMed:22699938, PubMed:29739711). Interacts with RORA. May interact with P-cadherin/CDH3. Interacts with RNF220 (PubMed:25266658). Interacts with CTNND2 (PubMed:25807484). Interacts (via the C-terminal region) with CBY1 (PubMed:12712206, PubMed:16424001). The complex composed, at least, of APC, CTNNB1 and GSK3B interacts with JPT1; the interaction requires the inactive form of GSK3B (phosphorylated at 'Ser-9') (PubMed:25169422). Interacts with DLG5 (By similarity). Interacts with FAM53B; promoting translocation to the nucleus (PubMed:25183871). Interacts with TMEM170B (PubMed:29367600). Interacts with AHI1 (PubMed:21623382). Interacts with GID8 (PubMed:28829046). Component of an cadherin:catenin adhesion complex composed of at least of CDH26, beta-catenin/CTNNB1, alpha-catenin/CTNNA1 and p120 catenin/CTNND1 (PubMed:28051089). Forms a complex comprising APPL1, RUVBL2, APPL2, HDAC1 and HDAC2 (PubMed:19433865). Interacts with IRF2BPL; mediates the ubiquitination and degradation of CTNNB1 (PubMed:29374064). Interacts with AMFR (By similarity). Interacts with LMBR1L (PubMed:31073040). Interacts with SOX30; prevents interaction of CTNNB1 with TCF7L2/TCF4 and leads to inhibition of Wnt signaling (PubMed:29739711). Interacts with SOX9; inhibiting CTNNB1 activity by competing with the binding sites of TCF/LEF within CTNNB1, thereby inhibiting the Wnt signaling (By similarity). Interacts with SPN/CD43 cytoplasmic tail (PubMed:15003504). Interacts (when phosphorylated at Tyr-333) with isoform M2 of PKM (PKM2); promoting transcription activation (PubMed:22056988). Interacts with PKP2 (via HEAD domain) (PubMed:11790773). Interacts with CDH1 (PubMed:11790773). Interacts (when unphosphorylated) with FLYWCH1, perhaps preventing interaction of CTNNB1 with TCF4, and thereby regulating transcription activation; phosphorylation of CTNNB1 may inhibit the interaction (PubMed:30097457). Interacts (via the central armadillo domains) with probable transcriptional regulator ADNP (via N-terminal region); interaction is direct and stabilizes CTNNB1 by modulating its phosphorylation by glycogen synthase kinase-3 beta GSK3B (By similarity). Interacts with NR5A2 (PubMed:22187462). Interacts with DSG2; the interaction promotes localization of CTNNB1 at cell junctions thus reducing its nuclear localization and subsequent transcription of CTNNB1/TCF-target genes (PubMed:29910125). {ECO:0000250|UniProtKB:Q02248, ECO:0000269|PubMed:10966653, ECO:0000269|PubMed:11389839, ECO:0000269|PubMed:11389840, ECO:0000269|PubMed:11590244, ECO:0000269|PubMed:11713475, ECO:0000269|PubMed:11713476, ECO:0000269|PubMed:11751639, ECO:0000269|PubMed:11790773, ECO:0000269|PubMed:12077367, ECO:0000269|PubMed:12297051, ECO:0000269|PubMed:12370829, ECO:0000269|PubMed:12408824, ECO:0000269|PubMed:12420223, ECO:0000269|PubMed:12501215, ECO:0000269|PubMed:12712206, ECO:0000269|PubMed:12830000, ECO:0000269|PubMed:14595118, ECO:0000269|PubMed:15003504, ECO:0000269|PubMed:16424001, ECO:0000269|PubMed:16858403, ECO:0000269|PubMed:17009320, ECO:0000269|PubMed:17047063, ECO:0000269|PubMed:17052462, ECO:0000269|PubMed:17289029, ECO:0000269|PubMed:17524503, ECO:0000269|PubMed:18077326, ECO:0000269|PubMed:18086858, ECO:0000269|PubMed:18378692, ECO:0000269|PubMed:18819930, ECO:0000269|PubMed:19433865, ECO:0000269|PubMed:19693690, ECO:0000269|PubMed:19816403, ECO:0000269|PubMed:19996314, ECO:0000269|PubMed:20026641, ECO:0000269|PubMed:20164195, ECO:0000269|PubMed:21132015, ECO:0000269|PubMed:21247902, ECO:0000269|PubMed:21262353, ECO:0000269|PubMed:21623382, ECO:0000269|PubMed:22056988, ECO:0000269|PubMed:22155184, ECO:0000269|PubMed:22187462, ECO:0000269|PubMed:22647378, ECO:0000269|PubMed:22699938, ECO:0000269|PubMed:25169422, ECO:0000269|PubMed:25183871, ECO:0000269|PubMed:25266658, ECO:0000269|PubMed:25807484, ECO:0000269|PubMed:28051089, ECO:0000269|PubMed:28829046, ECO:0000269|PubMed:29367600, ECO:0000269|PubMed:29374064, ECO:0000269|PubMed:29432739, ECO:0000269|PubMed:29739711, ECO:0000269|PubMed:29910125, ECO:0000269|PubMed:31073040, ECO:0000269|PubMed:7982500, ECO:0000305|PubMed:31073040}.; SUBUNIT: (Microbial infection) Interacts with herpes virus 8 protein vPK; this interaction inhibits the Wnt signaling pathway. {ECO:0000269|PubMed:29432739}. |
| Post-translational modification PTM: Phosphorylation at Ser-552 by AMPK promotes stabilization of the protein, enhancing TCF/LEF-mediated transcription (By similarity). Phosphorylation by GSK3B requires prior phosphorylation of Ser-45 by another kinase (PubMed:10966653, PubMed:12027456, PubMed:12051714). Phosphorylation proceeds then from Thr-41 to Ser-37 and Ser-33 (PubMed:12077367, PubMed:25169422). Phosphorylated by NEK2 (PubMed:18086858). EGF stimulates tyrosine phosphorylation (PubMed:10187801). Phosphorylated on Ser-33 and Ser-37 by HIPK2 and GSK3B, this phosphorylation triggers proteasomal degradation (PubMed:20307497). Phosphorylation on Ser-191 and Ser-246 by CDK5 (PubMed:17009320). Phosphorylation by CDK2 regulates insulin internalization (PubMed:21262353). Phosphorylation by PTK6 at Tyr-64, Tyr-142, Tyr-331 and/or Tyr-333 with the predominant site at Tyr-64 is not essential for inhibition of transcriptional activity (PubMed:20026641). Phosphorylation by SRC at Tyr-333 promotes interaction with isoform M2 of PKM (PKM2); promoting transcription activation (PubMed:22056988). {ECO:0000250|UniProtKB:Q02248, ECO:0000269|PubMed:10187801, ECO:0000269|PubMed:10966653, ECO:0000269|PubMed:12027456, ECO:0000269|PubMed:12051714, ECO:0000269|PubMed:12077367, ECO:0000269|PubMed:17009320, ECO:0000269|PubMed:18086858, ECO:0000269|PubMed:20026641, ECO:0000269|PubMed:20307497, ECO:0000269|PubMed:21262353, ECO:0000269|PubMed:22056988, ECO:0000269|PubMed:25169422}.; PTM: Ubiquitinated by the SCF(BTRC) E3 ligase complex when phosphorylated by GSK3B, leading to its degradation (PubMed:12077367). Ubiquitinated by a E3 ubiquitin ligase complex containing UBE2D1, SIAH1, CACYBP/SIP, SKP1, APC and TBL1X, leading to its subsequent proteasomal degradation (PubMed:11389839, PubMed:11389840, PubMed:20307497). Ubiquitinated and degraded following interaction with SOX9 (By similarity). Ubiquitinated via 'Lys-11'- and 'Lys-29'-linked ubiquitin chains by UBR5, leading to its stabilization (PubMed:21118991). {ECO:0000250|UniProtKB:Q02248, ECO:0000269|PubMed:11389839, ECO:0000269|PubMed:11389840, ECO:0000269|PubMed:12077367, ECO:0000269|PubMed:20307497, ECO:0000269|PubMed:21118991}.; PTM: S-nitrosylation at Cys-619 within adherens junctions promotes VEGF-induced, NO-dependent endothelial cell permeability by disrupting interaction with E-cadherin, thus mediating disassembly adherens junctions. {ECO:0000250|UniProtKB:Q02248}.; PTM: O-glycosylation at Ser-23 decreases nuclear localization and transcriptional activity, and increases localization to the plasma membrane and interaction with E-cadherin CDH1. {ECO:0000269|PubMed:24342833}.; PTM: Deacetylated at Lys-49 by SIRT1. {ECO:0000269|PubMed:24824780}.; PTM: Phosphorylated at Thr-556 by herpes virus 1/HHV-1 leading to CTNNB1 inhibition. {ECO:0000269|PubMed:24824780}. |
| Involvement in disease DISEASE: Colorectal cancer (CRC) [MIM:114500]: A complex disease characterized by malignant lesions arising from the inner wall of the large intestine (the colon) and the rectum. Genetic alterations are often associated with progression from premalignant lesion (adenoma) to invasive adenocarcinoma. Risk factors for cancer of the colon and rectum include colon polyps, long-standing ulcerative colitis, and genetic family history. {ECO:0000269|PubMed:9065402}. Note=The gene represented in this entry may be involved in disease pathogenesis.; DISEASE: Note=Activating mutations in CTNNB1 have oncogenic activity resulting in tumor development. Somatic mutations are found in various tumor types, including colon cancers, ovarian and prostate carcinomas, hepatoblastoma (HB), hepatocellular carcinoma (HCC). HBs are malignant embryonal tumors mainly affecting young children in the first three years of life.; DISEASE: Pilomatrixoma (PTR) [MIM:132600]: Common benign skin tumor. {ECO:0000269|PubMed:10192393, ECO:0000269|PubMed:11703283, ECO:0000269|PubMed:12027456}. Note=The gene represented in this entry is involved in disease pathogenesis.; DISEASE: Medulloblastoma (MDB) [MIM:155255]: Malignant, invasive embryonal tumor of the cerebellum with a preferential manifestation in children. {ECO:0000269|PubMed:10666372, ECO:0000269|PubMed:12027456}. Note=The gene represented in this entry may be involved in disease pathogenesis.; DISEASE: Ovarian cancer (OC) [MIM:167000]: The term ovarian cancer defines malignancies originating from ovarian tissue. Although many histologic types of ovarian tumors have been described, epithelial ovarian carcinoma is the most common form. Ovarian cancers are often asymptomatic and the recognized signs and symptoms, even of late-stage disease, are vague. Consequently, most patients are diagnosed with advanced disease. {ECO:0000269|PubMed:10391090}. Note=Disease susceptibility is associated with variants affecting the gene represented in this entry.; DISEASE: Note=A chromosomal aberration involving CTNNB1 is found in salivary gland pleiomorphic adenomas, the most common benign epithelial tumors of the salivary gland. Translocation t(3;8)(p21;q12) with PLAG1. {ECO:0000269|PubMed:10029085, ECO:0000269|PubMed:9020842}.; DISEASE: Mesothelioma, malignant (MESOM) [MIM:156240]: An aggressive neoplasm of the serosal lining of the chest. It appears as broad sheets of cells, with some regions containing spindle-shaped, sarcoma-like cells and other regions showing adenomatous patterns. Pleural mesotheliomas have been linked to exposure to asbestos. {ECO:0000269|PubMed:11464291}. Note=The gene represented in this entry may be involved in disease pathogenesis.; DISEASE: Neurodevelopmental disorder with spastic diplegia and visual defects (NEDSDV) [MIM:615075]: An autosomal dominant disorder characterized by global developmental delay, severe intellectual disability with absent or very limited speech, microcephaly, spasticity, and visual abnormalities. {ECO:0000269|PubMed:23033978, ECO:0000269|PubMed:25326669, ECO:0000269|PubMed:28514307}. Note=The disease is caused by variants affecting the gene represented in this entry.; DISEASE: Vitreoretinopathy, exudative 7 (EVR7) [MIM:617572]: A form of exudative vitreoretinopathy, a disorder of the retinal vasculature characterized by an abrupt cessation of growth of peripheral capillaries, leading to an avascular peripheral retina. This may lead to compensatory retinal neovascularization, which is thought to be induced by hypoxia from the initial avascular insult. New vessels are prone to leakage and rupture causing exudates and bleeding, followed by scarring, retinal detachment and blindness. Clinical features can be highly variable, even within the same family. Patients with mild forms of the disease are asymptomatic, and their only disease related abnormality is an arc of avascular retina in the extreme temporal periphery. {ECO:0000269|PubMed:28575650}. Note=The disease is caused by variants affecting the gene represented in this entry. |
| Target Relevance information above includes information from UniProt accession: P35222 |
| The UniProt Consortium |
Data
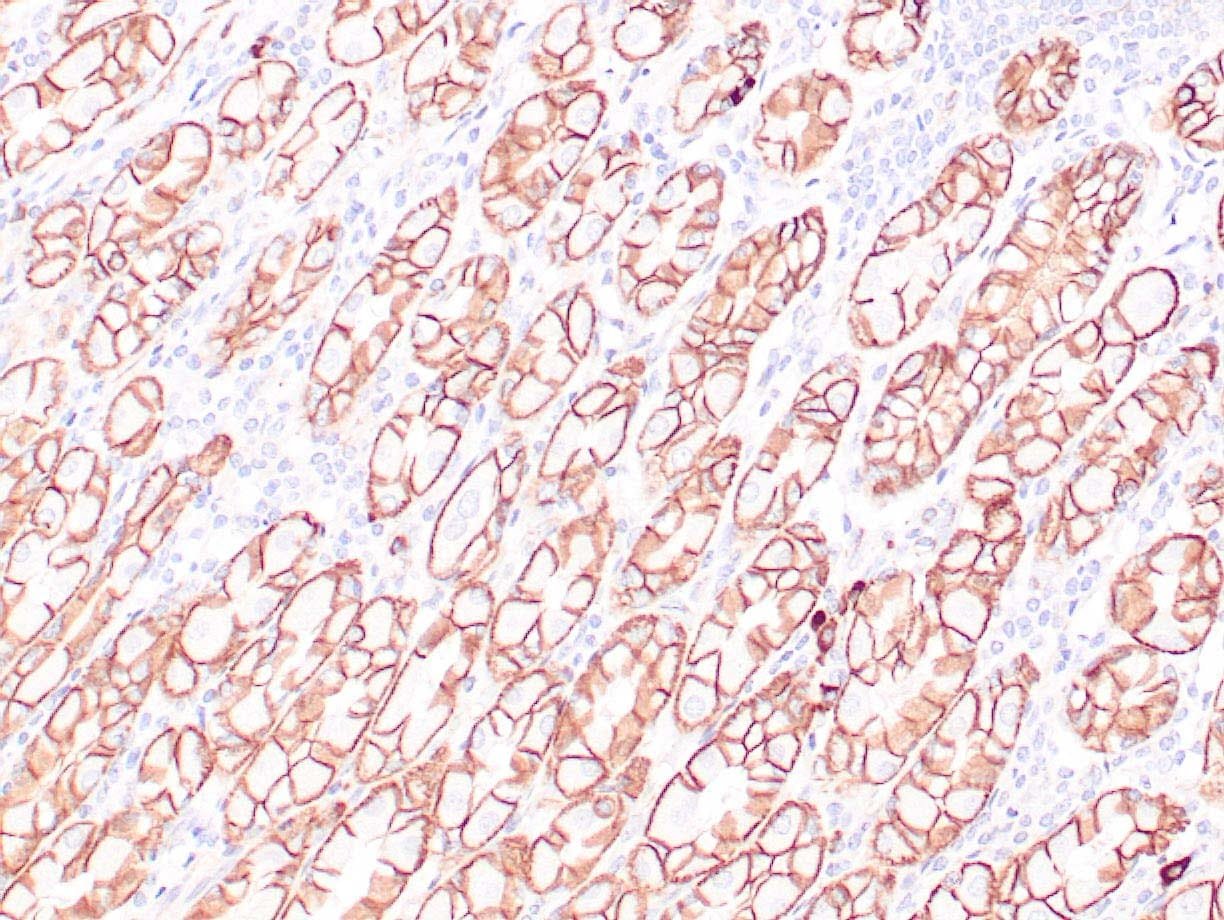 |
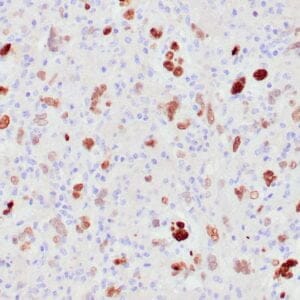
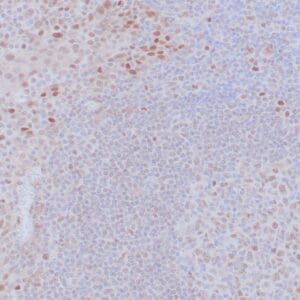
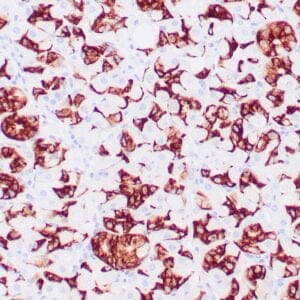
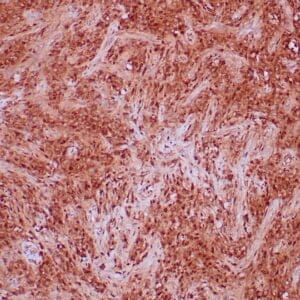
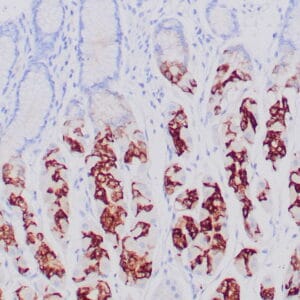
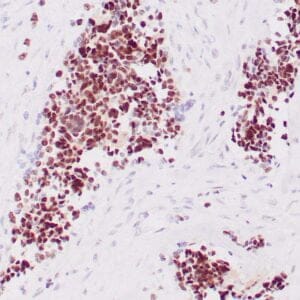
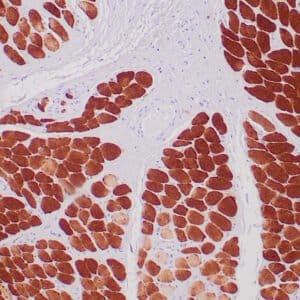
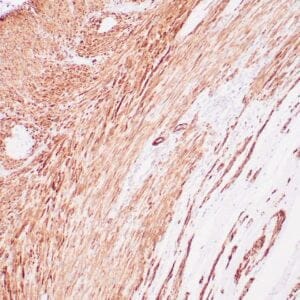
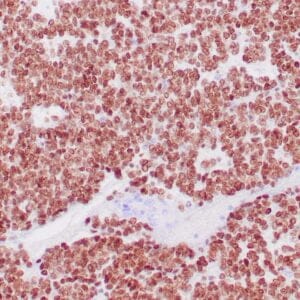
| Human colon adenocarcinoma stained with anti-beta-catenin antibody using peroxidase-conjugate and DAB chromogen. Note membranous staining of tumor cells. |
Publications
| pmid | title | authors | citation |
|---|---|---|---|
| We haven't added any publications to our database yet. | |||
Protocols
| relevant to this product |
|---|
| IHC |
Documents
| # | SDS | Certificate | |
|---|---|---|---|
| Please enter your product and batch number here to retrieve product datasheet, SDS, and QC information. | |||
Only logged in customers who have purchased this product may leave a review.
Reviews
There are no reviews yet.